New Design Rules for Ribosomal Monomers
Synthetic biology researchers created new substrates to guide the manufacture of new classes of polymers
Synthetic biologists at Northwestern Engineering have developed a set of design rules that guide how ribosomes can incorporate new kinds of monomers — a step forward in using these cellular machines to create new materials and therapeutics.
Recombinant protein production by the ribosome has transformed the lives of millions of people through the synthesis of biopharmaceuticals, like insulin, and industrial enzymes that are used in laundry detergents. In nature, however, the ribosome only incorporates natural amino acid monomers into protein polymers.
 “We set out to expand the range of ribosomal monomers for protein synthesis to enable new directions in biomanufacturing,” said Michael Jewett, the Charles Deering McCormick Professor of Teaching Excellence, professor of chemical and biological engineering, and director of the Center for Synthetic Biology at Northwestern’s McCormick School of Engineering. “What’s so exciting is that we learned the ribosome can accommodate more kinds of monomers than we expected, which sets the stage for using the ribosome as a general machine to create classes of materials and medicines that haven’t been synthesized before.”
“We set out to expand the range of ribosomal monomers for protein synthesis to enable new directions in biomanufacturing,” said Michael Jewett, the Charles Deering McCormick Professor of Teaching Excellence, professor of chemical and biological engineering, and director of the Center for Synthetic Biology at Northwestern’s McCormick School of Engineering. “What’s so exciting is that we learned the ribosome can accommodate more kinds of monomers than we expected, which sets the stage for using the ribosome as a general machine to create classes of materials and medicines that haven’t been synthesized before.”
Jewett was co-author of the research, which was published November 8 in the journal Nature Communications.
To expand the repertoire of monomers used by the ribosome, Jewett’s team set-out to identify design rules for linking monomers to transfer RNAs (tRNAs). That’s because getting the ribosome to use a new monomer is not as simple as introducing a new monomer to the ribosome. The monomers must be attached to tRNAs, which are the molecules that carry them into the ribosome. Many current processes for attaching monomers to tRNAs are difficult and time-consuming, but a relatively new process called flexizyme enables easier and more flexible attachment of monomers.
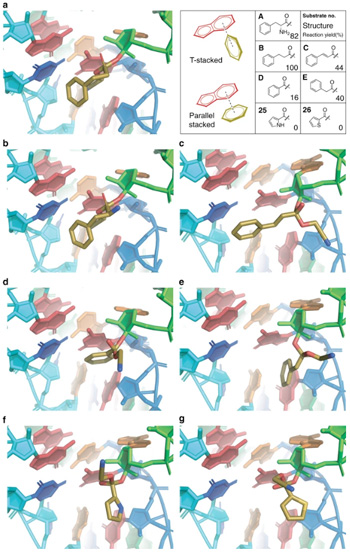 To develop the design rules for using flexizyme, the researchers created 37 monomers that were new to the ribosome from a diverse repertoire of scaffolds. Then, they showed that the monomers that could be attached to tRNAs could be used to make tens of new peptide hybrids. Finally, they validated their design rules by predictably guiding the search for even more new monomers.
To develop the design rules for using flexizyme, the researchers created 37 monomers that were new to the ribosome from a diverse repertoire of scaffolds. Then, they showed that the monomers that could be attached to tRNAs could be used to make tens of new peptide hybrids. Finally, they validated their design rules by predictably guiding the search for even more new monomers.
“With the new design rules, we show that we can avoid the trial-and-error approaches that have been historically associated with developing new monomers for use by the ribosome,” Jewett said.
These new design rules should accelerate the pace in which researchers can incorporate new monomers, which ultimately will lead to new bioproducts synthesized by the ribosome. For example, materials made of protease-resistant monomers could lead to antimicrobial drugs that combat rising antibiotic resistance.
The research is part of the Department of Defense’s Multidisciplinary University Research Initiatives (MURI) program, supported by the Army Research Office, in which Jewett is working with researchers from three other universities to reengineer the ribosome as a biological catalyst to make novel chemical polymers.
“These findings are an exciting step forward to achieving sequence-defined synthetic polymers which has been a grand challenge in the field of polymer chemistry,” said Dawanne Poree, lead program manager on this project at the Army Research Office. “The ability to harness and adapt cellular machinery to produce non-biological polymers would, in essence, bring synthetic materials into the realm of biological functions. This could render advanced, high-performance materials such as nanoelectronics, self-healing materials, and other materials of interest for the Army.”
The Army Research Office is an element of the US Army Combat Capabilities Development Command’s Army Research Laboratory.
“It’s amazing that the ribosome can accommodate the breadth of monomers we showed,” Jewett said. “That’s really encouraging for future efforts to repurpose ribosomes.”