New Discovery Makes It Easier to Design Synthetic Proteins that Rival their Natural Counterparts
The findings could have applications in water desalination, batteries, and pharmaceutical and biofuels research
A multi-institutional research team including Northwestern Engineering’s Monica Olvera de la Cruz has created a synthetic material that is as effective as naturally occurring proteins in transporting molecules through membranes, a major milestone that could transform such fields as medicine, life sciences, alternative energy, and environmental science.
Mimicking transmembrane proteins, which act as gatekeepers in living cells, has been a key goal — and a significant bottleneck — in synthetic membrane development for applications such as water desalination, batteries, and pharmaceutical and biofuels research.
This new achievement, described in a study published January 8 in the journal Nature, could alleviate that jam.
 “It’s surprising that synthetic random polymers, or a collection of polymers with various sequences along their backbone, can selectively aid proton transport in cell membranes,” said Olvera de la Cruz, Lawyer Taylor Professor of Materials Science and Engineering at the McCormick School of Engineering and a co-author of the study. “Our goal was to find random polymers that would mimic the job of intrinsically disordered proteins in cell media, which can concentrate enzymes to control chemical reaction rates. We ended up finding that these random polymers can also be adsorbed in the cell membrane, suggesting that intrinsically disordered proteins can also act as transmembrane proteins.”
“It’s surprising that synthetic random polymers, or a collection of polymers with various sequences along their backbone, can selectively aid proton transport in cell membranes,” said Olvera de la Cruz, Lawyer Taylor Professor of Materials Science and Engineering at the McCormick School of Engineering and a co-author of the study. “Our goal was to find random polymers that would mimic the job of intrinsically disordered proteins in cell media, which can concentrate enzymes to control chemical reaction rates. We ended up finding that these random polymers can also be adsorbed in the cell membrane, suggesting that intrinsically disordered proteins can also act as transmembrane proteins.”
“The challenge is that there exists a large number of possible sequences for random polymers,” said Baofu Qiao, research assistant professor in Northwestern Engineering’s Center for Computation and Theory of Soft Materials and study co-author. “Amazingly, nature picks its favorite in orientating the amphiphilic random polymers so that the nonpolar monomers go into the interior of lipid membranes with the polar monomers protruding into water. These membrane polymers are structurally and functionally similar to transmembrane proteins, providing a new opportunity for bio-inspired materials.”
This research builds on previous work by the researchers in which random polymers were used as synthetic chaperones to stabilize enzymes in non-natural environments. In that study, published in Science in 2018, Olvera de la Cruz and Qiao used molecular simulations to show that the polymers interacted favorably with protein surfaces by wrapping around protein surfaces in organic solvents and adhering weakly in water, leading to correct protein folding and stability in a non-native environment. A follow-up study expanded the work by further exploring the resemblance between random polymers and intrinsically disordered proteins.
“We showed then that the composition of random polymers can be designed to cover many different enzymes in different solvents,” Olvera de la Cruz said. “We determined the optimal composition of hydrophobic and hydrophilic groups that work to protect and deliver to different media enzymes that have hydrophobic and hydrophilic groups on their surfaces.”
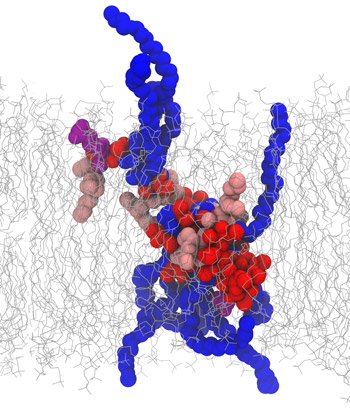 In this new paper, Olvera de la Cruz, Qiao, and collaborators from the University of California at Berkeley, UC Santa Cruz, Oak Ridge National Laboratory designed a polymer that selectively transported protons — charged subatomic particles — across an acrylic film at a rate similar to those of natural proton channels while successfully filtering out other types of cations.
In this new paper, Olvera de la Cruz, Qiao, and collaborators from the University of California at Berkeley, UC Santa Cruz, Oak Ridge National Laboratory designed a polymer that selectively transported protons — charged subatomic particles — across an acrylic film at a rate similar to those of natural proton channels while successfully filtering out other types of cations.
Selective and rapid proton transport is important in the regulation of pH levels and many biological functions. That capability is also useful for clean energy such as fuel cells and energy storage.
“If we think of the polymer as a doorway, it had to be open wide enough for the proton to go through, but not so open that other cations can sneak in,” said Ting Xu, professor of materials science and engineering and chemistry at Berkeley, who led the study. “We were able to find that right balance with our material.”
The paper also provided a solution to a long-standing challenge in designing synthetic proteins that worked like their natural counterparts. Monomer molecules bond together to form polymers, and amino acids are monomers that form proteins. The monomer sequence governs protein function, so for decades scientists believed that they had to copy the monomer sequence exactly to create a synthetic polymer that would function as well as naturally occurring proteins.
Previous attempts to fabricate synthetic protein channels have used sequence-defined polymers — as well as peptides and carbon nanotubes — but none rivaled the performance of their natural counterparts.
Xu and her collaborators found that the monomers in the polymer don’t have to line up nor match exactly to function like a protein. Rather, their design only requires researchers to statistically control the sequence of four types of monomers — Xu calls them random heteropolymers (RHPs) — for the polymer to perform as well as a naturally occurring protein. The monomers can be grouped into segments like Lego pieces to construct functional protein-mimics.
“Compare this to how cars are built,” Xu said. “There are different models, colors, and shapes, but they all contain important parts such as an engine, wheels, and energy source. For each part, there can be different options, such as gas or electric engines, but at the end of the day, it’s a car, not a train.”
Xu and her team designed a library of polymers that are statistically similar in sequence, providing newfound flexibility in assembly.
“What makes our new technique promising is that it’s scalable, and the knowledge to do this is readily available,” Xu said. “Considering the vast number of monomers available and the recent advances in polymer chemistry, the possibilities of marrying the synthetic and biological fields are almost unlimited. Nature is simple and thrifty. It is our job to crack its code to live in harmony.”
Other co-authors on the paper include Tao Jiang, Aaron Hall, Marco Eres, Yun Zhou, Zhiyuan Ruan, Andrew Couse, and Haiyan Huang of UC Berkeley, Zahra Hemmatian and Marco Rolandi of UC Santa Cruz, and William Heller of Oak Ridge National Laboratory.
This work was supported by the Army Research Office and the National Science Foundation.